Clustered placement gallery (analytic vs. bootstrap)
This short gallery compares analytic and bootstrap DBH payloads under the clustered placement mode.
It reuses the seeded fixture tests/fixtures/synthesis/clustered_gallery.json so plots stay
reproducible across refactors.
Load the fixture and compute summary stats
from pathlib import Path
import json
import numpy as np
payload = json.loads(Path("tests/fixtures/synthesis/clustered_gallery.json").read_text())
polygon = np.asarray(payload["polygon"], dtype=float)
print(payload["analytic"]["mean_dbh"], payload["bootstrap"]["mean_dbh"])
Expected means (seed=2025, cluster_spread=0.08, min_spacing=0.05, 2×1.5 polygon):
Analytic (lognormal μ=2.0, σ²=0.25, n=10): mean DBH ≈ 11.56 cm (σ ≈ 6.79).
Bootstrap (empirical vectors, n=10): mean DBH ≈ 14.20 cm (σ ≈ 2.97).
Visualise clustered placement
import matplotlib.pyplot as plt
from nemora.synthesis import stands, stems
def analytic_sampler():
entry = stands.StandBootstrapLibraryEntry(
identifier="analytic-1",
source="analytic",
metadata={"distribution": "lognormal", "parameters": {"mu": 2.0, "sigma2": 0.25}, "sample_size": 10},
dbh_vectors={},
)
assignment = stands.StandBootstrapAssignment(
stand_id="stand-0001",
vegetation_type="fir",
age_class="60-80",
area=4.0,
bootstrap_id="analytic-1",
)
return stems.StandDBHSampler(assignment=assignment, entry=entry)
def bootstrap_sampler(vectors):
entry = stands.StandBootstrapLibraryEntry(
identifier="bootstrap-1",
source="bootstrap.json",
metadata={"distribution": "empirical", "sample_size": 10, "mode": "bootstrap"},
dbh_vectors=vectors,
)
assignment = stands.StandBootstrapAssignment(
stand_id="stand-0002",
vegetation_type="pine",
age_class="40-60",
area=3.5,
bootstrap_id="bootstrap-1",
)
return stems.StandDBHSampler(assignment=assignment, entry=entry)
fixture = json.loads(Path("tests/fixtures/synthesis/clustered_gallery.json").read_text())
polygon = np.asarray(fixture["polygon"], dtype=float)
rng = np.random.default_rng(fixture["analytic"]["seed"])
anal_records = stems.place_trees_with_dbh(
polygon,
analytic_sampler(),
rng=rng,
config=stems.TreePlacementConfig(mode="clustered", cluster_spread=fixture["analytic"]["cluster_spread"], min_spacing=0.05),
)
rng = np.random.default_rng(fixture["bootstrap"]["seed"])
boot_records = stems.place_trees_with_dbh(
polygon,
bootstrap_sampler(fixture["bootstrap"]["vectors"]),
rng=rng,
config=stems.TreePlacementConfig(mode="clustered", cluster_spread=fixture["bootstrap"]["cluster_spread"], min_spacing=0.05),
)
fig, ax = plt.subplots(figsize=(6, 4))
poly = np.vstack([polygon, polygon[0]]) # close ring for plotting
ax.plot(poly[:, 0], poly[:, 1], color="black", lw=1, label="polygon")
ax.scatter(
[r["x"] for r in anal_records],
[r["y"] for r in anal_records],
c=[r["dbh"] for r in anal_records],
cmap="Blues",
marker="o",
label="analytic",
alpha=0.8,
)
ax.scatter(
[r["x"] for r in boot_records],
[r["y"] for r in boot_records],
c=[r["dbh"] for r in boot_records],
cmap="Oranges",
marker="x",
label="bootstrap",
alpha=0.9,
)
ax.set_title("Clustered placement: analytic vs. bootstrap DBH")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
ax.set_aspect("equal")
plt.tight_layout()
plt.savefig("docs/examples/clustered_gallery.png", dpi=200)
Rendered preview (placeholder coefficients):
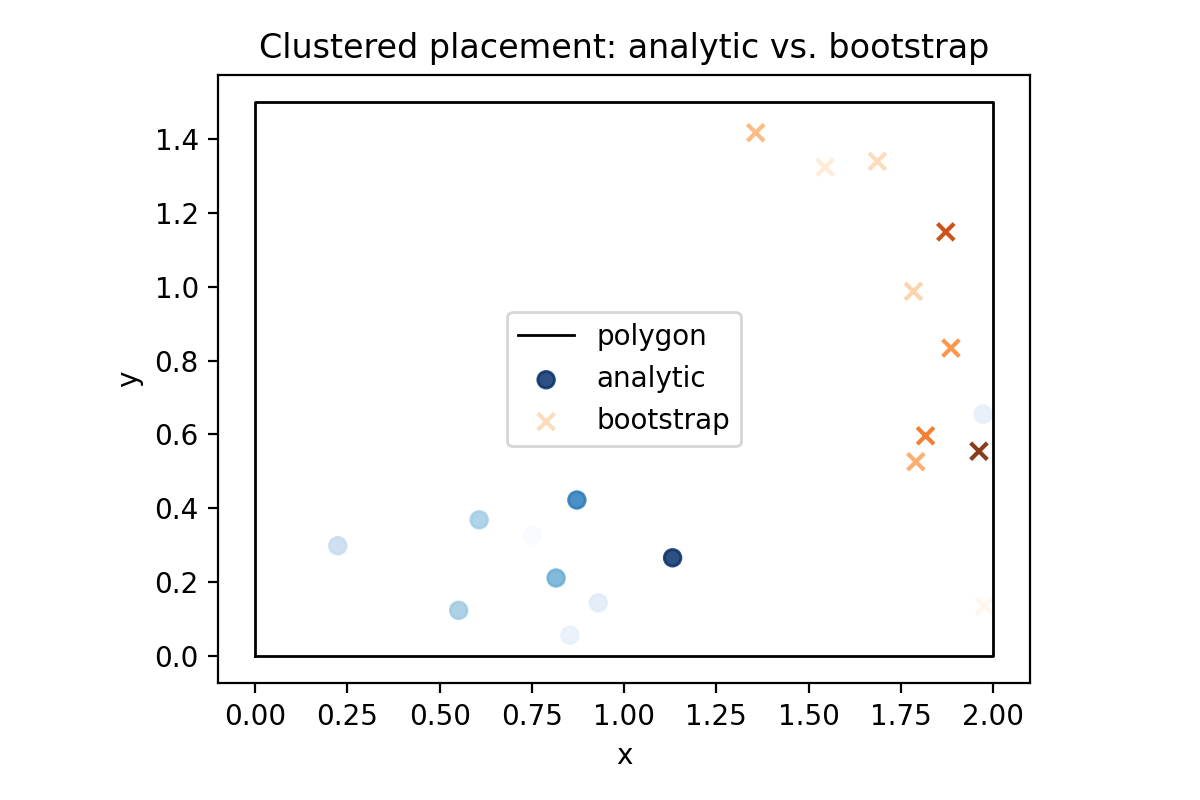
If placement or DBH draws change, rerun the snippet and compare against the expected means from the
fixture; the regression tests (`tests/test_synthesis_stems.py::test_clustered_gallery_fixture_alignment`)
will also fail if behaviour drifts.