Fit HPS Inventories
This guide walks through fitting the horizontal point sampling (HPS) workflow using the
nemora CLI and Python API. It extends the weighted estimator described in the UBC FRESH Lab
manuscript and relies on the probability distributions catalogued in the
distribution reference.
Prerequisites
An HPS tally file with columns
dbh_cm(bin midpoints) andtally(per-plot counts).The basal area factor (
BAF) used during cruise design.Python environment with
nemorainstalled. One approach:
python -m venv .venv
source .venv/bin/activate
pip install -e ".[dev]"
CLI Workflow
Inspect available distributions:
nemora registry
Fit the HPS inventory with the default candidate set (
weibull,gamma):
nemora fit-hps data/hps_tally.csv --baf 2.0
Pass additional --distribution options to try alternative PDFs as the CLI evolves. The command
prints RSS, small-sample AIC (AICc), Pearson chi-square, Kolmogorov–Smirnov, Cramér–von Mises, and
Anderson–Darling statistics (plus residual summaries) for each candidate distribution so you can
compare fits at a glance.
Generalised secant mixtures are exposed as gsmN (for any integer N ≥ 2), so a command such as
nemora fit-hps ... -d gsm3 -d gsm6 will evaluate both three- and six-component blends.
Use --distribution (repeatable) to limit the candidate list and --show-parameters to include the
fitted parameter estimates. For example:
nemora fit-hps data/hps_tally.csv --baf 2.0 -d weibull -d gamma --show-parameters
The grouped Weibull solver defaults to auto, which refines the least-squares seed with a Newton
update and quietly falls back to the stable least-squares solution if the grouped maximum-likelihood
step drifts. Use --grouped-weibull-mode ls to pin the legacy behaviour or
--grouped-weibull-mode mle to require the grouped likelihood path (the command will exit with an
error if the refinement fails under the forced mle mode).
To stay numerically stable, the solver applies the conditional offset from the manuscript:
the Weibull support is shifted by min(DBH) – 0.5 cm before evaluating grouped probabilities.
Johnson SB fits now run an EM pass that updates the Beta shape parameters in grouped space, while
Birnbaum–Saunders grouped fits attempt a truncated-normal EM step before falling back to the legacy
grouped MLE. Diagnostics include the solver mode (grouped-em or grouped-mle) so you can trace
which path executed.
To explore the public PSP example bundle prepared with
scripts/prepare_hps_dataset.py, start with one of the manifests in
examples/hps_baf12:
nemora fit-hps examples/hps_baf12/4000002_PSP1_v1_p1.csv --baf 12
Python API Example
import pandas as pd
from nemora.workflows import fit_hps_inventory
data = pd.read_csv("data/hps_tally.csv")
results = fit_hps_inventory(
dbh_cm=data["dbh_cm"].to_numpy(),
tally=data["tally"].to_numpy(),
baf=2.0,
distributions=["weibull", "gamma", "gb2"],
)
best = min(results, key=lambda r: r.gof["rss"])
print(best.distribution, best.parameters)
for result in results:
gof = result.gof
summary = result.diagnostics.get("residual_summary", {})
print(
f"{result.distribution:8s} RSS={gof.get('rss', float('nan')):,.2f} "
f"AICc={gof.get('aicc', float('nan')):,.2f} Chi^2={gof.get('chisq', float('nan')):,.2f} "
f"Max|res|={summary.get('max_abs', float('nan')):.3f}"
)
fit_hps_inventory expands tallies to stand tables, applies the HPS compression factors as
weights, and auto-generates starting values. Override the defaults through FitConfig for
specialised scenarios, and pass grouped_weibull_mode="ls" or "mle" to mirror the CLI toggle
when you need to pin or force the grouped Weibull solver mode.
Reference Fit (BC PSP 4000002-PSP1)
Using the public bundle prepared in the HPS dataset guide, fit_hps_inventory
identifies the Weibull distribution as the best fit for plot 4000002_PSP1_v1_p1
with BAF 12. This mirrors the methodology from the EarthArXiv preprint of the HPS
manuscript (Paradis, 2025). The regression test in tests/test_hps_parity.py
locks these targets:
Metric |
Value |
|---|---|
Distribution |
|
RSS |
|
Parameters |
|
Re-run the check locally with:
pytest tests/test_hps_parity.py
Censored Meta-Plot Demo
For a pooled view, the notebook examples/hps_bc_psp_demo.ipynb aggregates the public BC PSP files,
censors stems below 9 cm, and fits the censored workflow. This serves as a deployment example on new
data rather than a reproduction of the manuscript figures. Run the notebook (or
pytest tests/test_censored_workflow.py) to verify the gamma fit parameters recorded in the summary
table.
Parity Summary
Regression tests
tests/test_hps_parity.pyandtests/test_censored_workflow.pylock the manuscript-aligned Weibull and censored gamma fits.tests/test_cli.py::test_fit_hps_command_outputs_weibull_firstexercises the Typer CLI against the PSP tallies to ensure the command-line workflow mirrors the Python API.examples/hps_bc_psp_demo.ipynbdemonstrates the workflow on the public BC PSP dataset, whileexamples/hps_parity_reference.ipynbruns the parity analysis on the manuscript meta-plot dataset included underexamples/data/reference_hps/binned_meta_plots.csv(see Figure 1). The notebook also exportsdocs/_static/reference_hps_parity_table.csvsummarising RSS, AICc, chi-square, and parameter deltas for each meta-plot.For a pure-Python walkthrough (no notebooks), see the programmatic HPS guide.
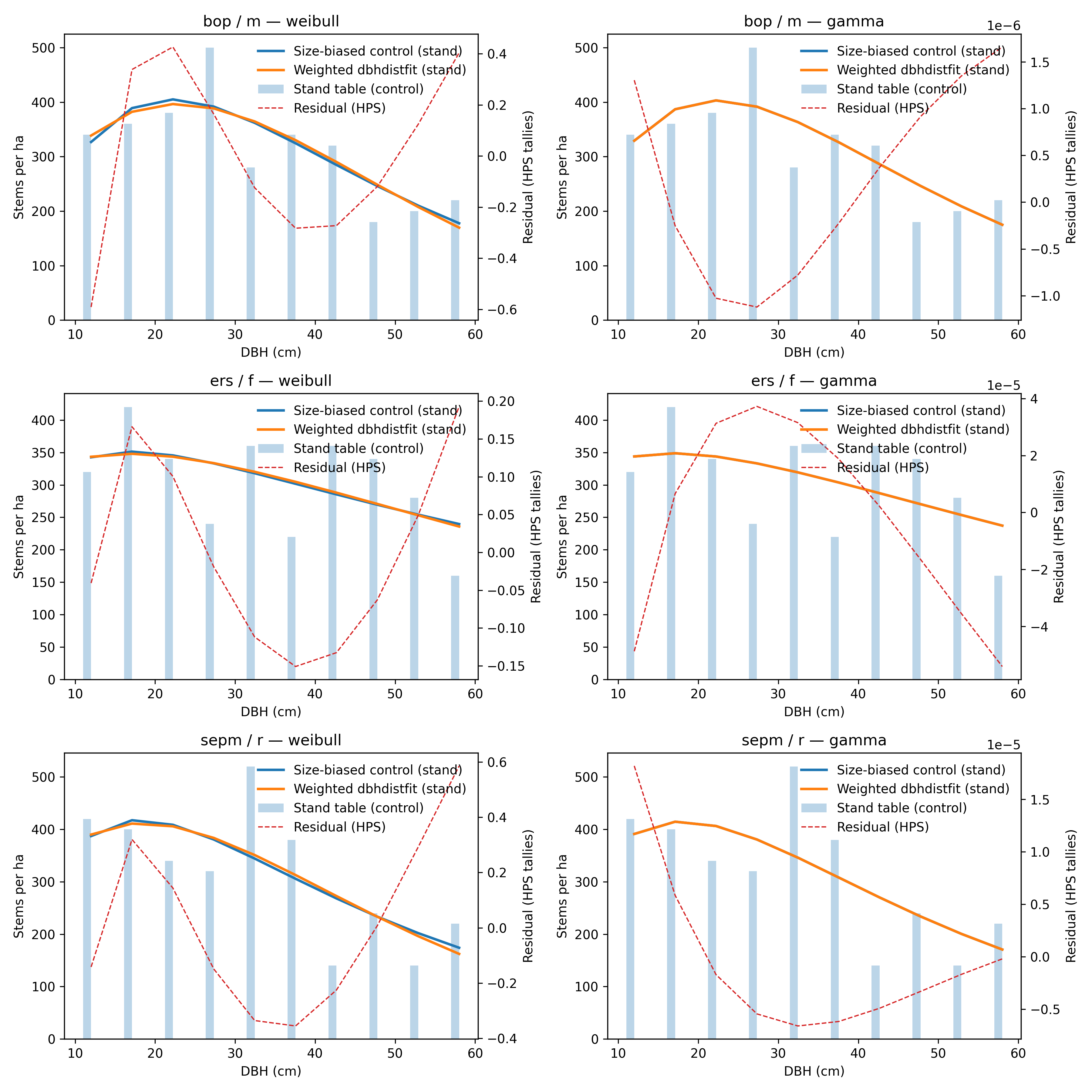
Figure 1 — Size-biased control vs. weighted nemora curves for the manuscript meta-plots. The
dashed line shows residuals on the HPS tally scale.
Censored / Two-Stage Baseline
The reference notebook also aggregates the manuscript dataset into a censored meta-plot (DBH ≥ 20 cm)
and fits the two-stage workflow via fit_censored_inventory. The exported table
(docs/_static/reference_hps_censored_table.csv) records RSS, AICc, chi-square, and parameter values
for each candidate distribution.
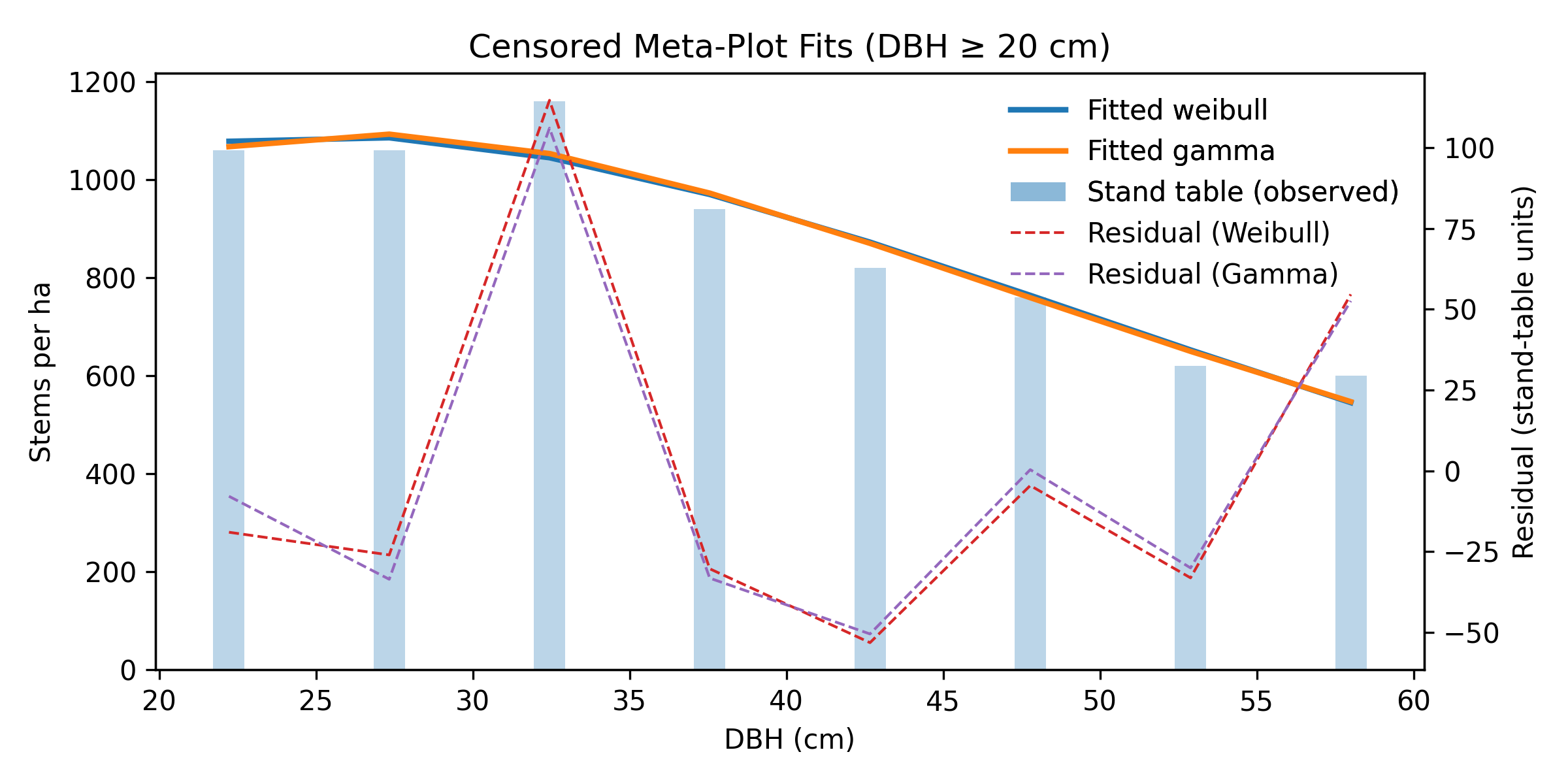
Figure 2 — Censored (DBH ≥ 20 cm) stand-table data with fitted Weibull and Gamma curves using the two-stage workflow. Residual lines highlight the agreement on the truncated support.
Diagnostics
Inspect
result.diagnostics["residuals"]for shape or bias.Compare
result.gofmetrics (RSS, AICc when available) across candidates.Plot the empirical stand table alongside fitted curves to confirm agreement with the manuscript workflow.
Next Steps
Expose distribution filters and parameter previews in the CLI.
Add worked examples for censored inventories and DataLad-backed datasets.
Integrate notebook tutorials mirroring the published reproducibility bundles.
Expand FAIR dataset coverage (see HPS dataset guide) with additional PSP plots and censored variants to support end-to-end parity tests.
.. todo:: Update this section once the nemora.ingest / sampling / synthesis modules land to reflect the broader workflow.